OSG fuels a student-developed computing platform to advance RNA nanomachines
Josephine Watkins August 10, 2021
How undergraduates at the University of Nebraska-Lincoln developed a science gateway that enables researchers to build RNA nanomachines for therapeutic, engineering, and basic science applications.

The UNL students involved in the capstone project, on graduation day. Order from left to right: Evan, Josh, Dan, Daniel, and Conner.
When a science gateway built by a group of undergraduate students is deployed this fall, it will open the door for researchers to leverage the capabilities of advanced software and the capacity of the Open Science Pool (OSPool). Working under the guidance of researcher Joe Yesselman and longtime OSG contributor Derek Weitzel, the students united advanced simulation technology and a national, open source of high throughput computing capacity –– all within an intuitive, web-accessible science gateway.
Joe, a biochemist, has been fascinated by computers and mathematical languages for as long as he can remember. Reminiscing to when he first adopted computer programming and coding as a hobby back in high school, he reflects: “English was difficult for me to learn, but for some reason mathematical languages make a lot of sense to me.”
Today, he is an Assistant Professor of Chemistry at the University of Nebraska-Lincoln (UNL), and his affinity for computer science hasn’t waned. Leading the Yesselman Lab, he relies on the interplay between computation and experimentation to study the unique structural properties of RNA.
In September of 2020, Joe began collaborating with UNL’s Holland Computing Center (HCC) and the OSG to accelerate RNA nanostructure research everywhere by making his lab’s RNAMake software suite accessible to other scientists through a web portal. RNAMake enables researchers to build nanomachines for therapeutic, engineering, and basic science applications by simulating the 3D design of RNA structures.
Five UNL undergraduate students undertook this project as part of a year-long computer science capstone experience. By the end of the academic year, the students developed a science gateway –– an intuitive web-accessible interface that makes RNAMake easier and faster to use. Once it’s deployed this fall, the science gateway will put the Yesselman Lab’s advanced software and the shared computing resources of the OSPool into the hands of researchers, all through a mouse and keyboard.
The gateway’s workflow is efficient and simple. Researchers upload their input files, set a few parameters, and click the submit button –– no command lines necessary. Short simulations will take merely a few seconds, while complex simulations can last up to an hour. Once the job is completed, an email appears in their inbox, prompting them to analyze and download the resulting RNA nanostructures through the gateway.
This was no small feat. Collaboration among several organizations brought this seemingly simple final product to fruition.
To begin the process, the students received a number of startup allocations from the Extreme Science and Engineering Discovery Environment (XSEDE). When it was time to build the application, they used Apache Airavata to power the science gateway and they extended this underlying software in some notable ways. In order to provide researchers with more intuitive results, they implemented a table viewer and a 3D molecule visualization tool. Additionally, they added the ability for Airavata to submit directly to HTCondor, making it possible for simulations to be distributed across the resources offered by the OSPool.
The simulations themselves are small, short, and can be run independently. Furthermore, many of these simulations are needed in order to discover the right RNA nanostructures for each researcher’s purpose. Combined, these qualities make the jobs a perfect candidate for the OSPool’s distributed high throughput computing capabilities, enabled by computing capacity from campuses across the country.
Commenting on the incorporation of OSG resources, project sponsor Derek Weitzel explains how the gateway “not only makes it easier to use RNAMake, but it also distributes the work on the OSPool so that researchers can run more RNAMake simulations at the same time.” If the scientific process is like a long road trip, using high throughput computing isn’t even like taking the highway –– it’s like skipping the road entirely and taking to the skies in a high-speed jet.
The science gateway has immense potential to transform the way in which RNA nanostructure research is conducted, and the collaboration required to build it has already made lasting impacts on those involved. The group of undergraduate students are, in fact, no longer undergraduates. The team’s student development manager, Daniel Shchur, is now a software design engineer at Communication System Solutions in Lincoln, Nebraska. Reflecting on the capstone project, he remarks, “I think the most useful thing that my teammates and I learned was just being able to collaborate with outside people. It was definitely something that wasn’t taught in any of our classes and I think that was the most invaluable thing we learned.”
But learning isn’t just exclusive to students. Joe notes that he gained some unexpected knowledge from the students and Derek. “I learned a ton about software development, which I’m actually using in my lab,” he explains. “It’s very interesting how people can be so siloed. Something that’s so obvious, almost trivial for Derek is something that I don’t even know about because I don’t have that expertise. I loved that collaboration and I loved hearing his advice.”
In the end, this collaboration vastly improved the accessibility of RNAMake, Joe’s software suite and the focus of the science gateway. Perhaps he explains it best with an analogy: ”RNAMake is basically a set of 500 different LEGO® pieces. Using enthusiastic gestures, Joe continues by offering an example: “Suppose you want to build something from this palm to this palm, in three-dimensional space. It [RNAMake] will find a set of LEGO® pieces that will fit there.”
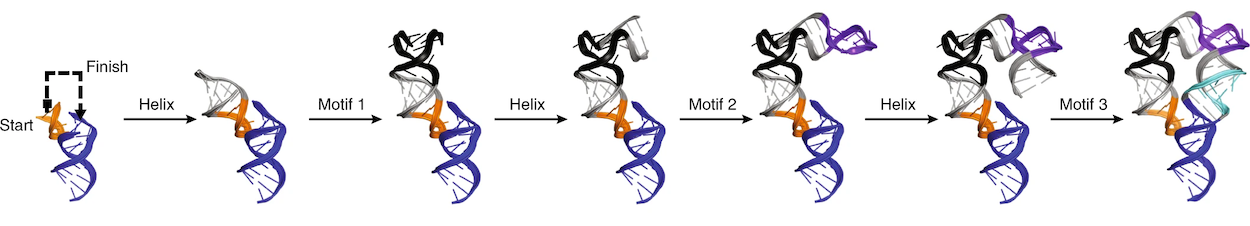
A demonstration of how RNAMake’s design algorithm works. Credit: Yesselman, J.D., Eiler, D., Carlson, E.D. et al. Computational design of three-dimensional RNA structure and function. Nat. Nanotechnol. 14, 866–873 (2019). https://doi.org/10.1038/s41565-019-0517-8
Since the possible combinations of these LEGO® pieces of RNA are endless, this tool saves users the painstaking work of predicting the structures manually. However, the installation and use of RNAMake requires researchers to have a large amount of command line knowledge –– something that the average biochemist might not have.
Ultimately, the science gateway makes this previously complicated software suddenly more accessible, allowing researchers to easily, quickly, and accurately design RNA nanostructures.
These structures are the basis for RNA nanomachines, which have a vast range of applications in society. Whether it be silencing RNAs that are used in clinical trials to cut cancer genes, or RNA biosensors that effectively bind to small molecules in order to detect contaminants even at low concentrations –– the RNAMake science gateway can help researchers design and build these structures.
Perhaps the most relevant and pressing applications are RNA-based vaccines like Moderna and Pfizer. These vaccines continue to be shipped across cities, countries, and continents to reach people in need, and it’s crucial that they remain in a stable form throughout their journey. Insight from RNA nanostructures can help ensure that these long strands of mRNA maintain stability so that they can eventually make their way into our cells.
Looking to the future, a second science gateway capstone project is already being planned for next year at UNL. Although it’s currently unclear what field of research it will serve, there’s no doubt that this project will foster collaboration, empower students and researchers, and impact society –– all through a few strokes on a keyboard.
